Notice: This essay builds directly on the previous post and assumes some familiarity with the experiment described there.
7th April 26
Update: Image classification by evolving bytecode
We report further progress on a machine learning strategy we have presented previously: evolving Zyme bytecode programs to solve a subset of the MNIST image classification problem. Among a number of incremental refinements, the most notable is the incorporation of ‘junk’ bytes - inspired by the role of non-coding DNA in biological genomes - as a mechanism for improving evolvability. Together, these advances bring us to a performance milestone of ~75% accuracy. We discuss how this work represents a departure from the dominant paradigm in machine learning: rather than treating the problem as one of parameter estimation within predefined model structures, we frame it as one of algorithm discovery - the autonomous identification of sequences of instructions capable of solving a task.
1. Introduction
We are developing a novel machine learning approach that reimagines genetic programming, a family of methods based on the principles of evolution by natural selection. Like genetic programming broadly, our method generates computer programs capable of solving a given task by iteratively selecting, mutating, and recombining best-performing candidates from a population. We extend this foundation with Zyme, our evolvable programming language and virtual machine, built specifically to support open-ended algorithm discovery.
In a previous post, we described this approach in detail and demonstrated consistent accuracy improvements on a four-digit subset of the MNIST image classification dataset. At that stage, however, the best programs reached only around 50% accuracy - a promising start, but well short of anything that could be called competitive. Since then, we have continued refining the method, with a focus on both accuracy and computational efficiency, and have recently succeeded in evolving programs that reach ~75% accuracy.
Beyond the accuracy gain itself, these results are significant for how they were achieved. Rather than relying on parameter estimation within the confines of predefined model structures, we treat machine learning as a problem of algorithm discovery: the autonomous identification of computational procedures (sequences of byte-level operations) capable of solving a task, with no fixed architecture or prior assumptions about what form a solution should take. Here, the algorithms take the form of the bytecode programs we are evolving. This framing - machine learning as algorithm discovery - represents a reorientation in the prevailing paradigm of how artificial intelligence is pursued.
2. Results
To evolve computer programs capable of classifying images, we followed the same two step-process as in the previous experiments: (1) reproduction with random mutation followed by (2) selection, in which the top-performing programs are retained to form the population for the next generation (see Previous Methods 3.4). This time, we scaled the process by significantly increasing in training dataset size and the population parameters controlling how many programs are spawned each generation1.
We also changed the composition of the initial programs. As before, we used the Zyme compiler to generate the starting population (see Previous Methods 3.3). In earlier experiments, we seeded the population directly with raw compiler output: programs in their most minimal form, consisting only of the functional strands defined in the source code.
Here, we introduced ‘junk’ bytes into these initial programs. These are extra bytes that do not affect immediate program behavior, analogous to the non-coding DNA found in biological genomes2. We hypothesised that minimal programs are brittle: because nearly every byte encodes an active instruction, most mutations disrupt existing logic rather than expanding it. By adding ‘junk’ bytes, we provide space where point mutations (that change individual bytes) can effectively activate new instructions from inactive bytes, rather than overwriting functional code.
This ‘junk’ was introduced to programs in two ways. First, random no-op bytes were inserted into existing strands, increasing their length without altering their behaviour. Second, strands were duplicated with a high error rate, producing inactive copies that resembled the functional strands in structure but not in behaviour.
Using these junk-seeded populations, we ran two independent replicates for 100 generations. To measure generalization, we evaluated every program’s accuracy against the full test dataset at each generation (see Figure 1).
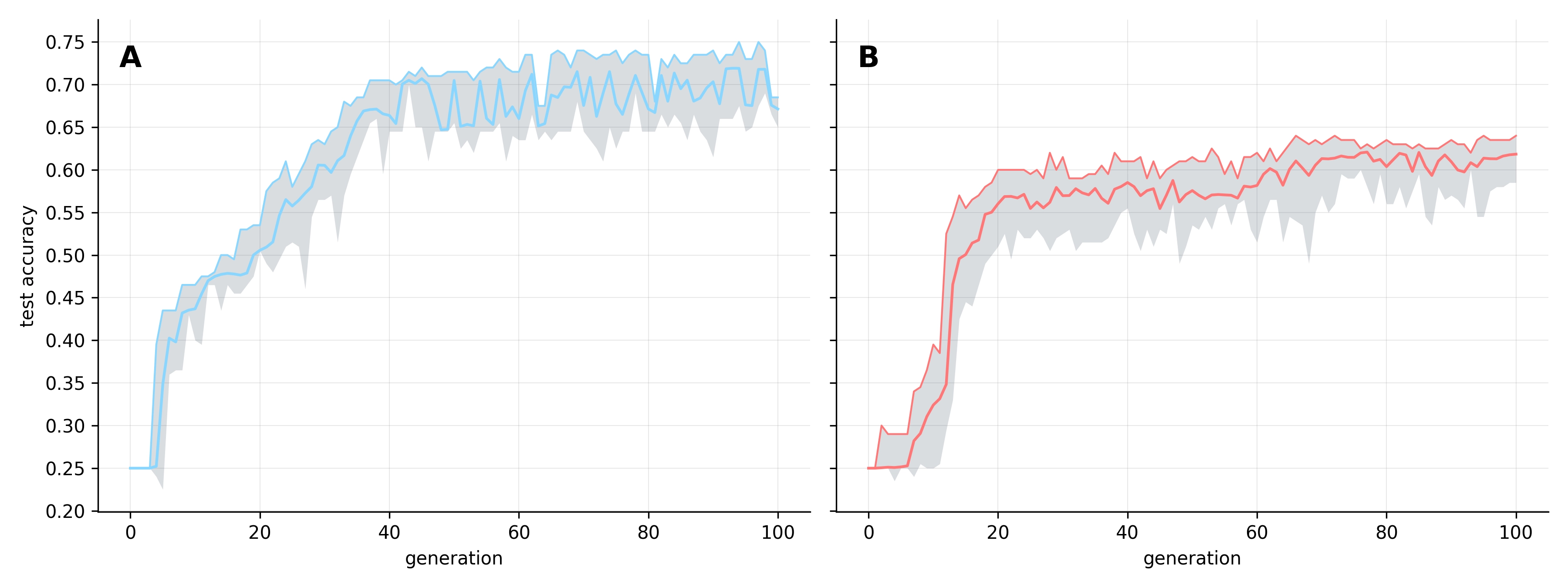
As observed in previous experiments, accuracy increases consistently across generations. However, these runs substantially exceeded our prior ceiling of ~50%, reaching mean population accuracies of ~73% and ~63% across the two replicates. Notably, individual programs in replicate A reached as high as 75% accuracy - a milestone that felt like a significant breakthrough.
The evolutionary trajectory in both replicates differed from our earlier experiments. Instead of the immediate gains seen previously, we observed a ‘lag phase’ of approximately five generations before accuracy improved meaningfully. In replicate B, while some individuals show above-random performance (>25% accuracy) from early on, the population as a whole does not begin to shift until around generation five; replicate A shows even less early activity. Once improvements do take hold, however, accuracy rises sharply and quickly surpasses the ceiling observed in previous experiments. Perhaps the 'junk' added to the initial programs makes it difficult to find early adaptive mutations, but paradoxically enhances long-term evolvability?
The two replicates converged on distinct peak accuracy scores, with Replicate A outperforming Replicate B by ~10%. This divergence reinforces our previous observation that independent runs often discover potentially entirely different mechanisms for solving the task. This difference is also visible in their stability: as replicate A approaches its peak, intergeneration variability increases as the population struggles to maintain its top-performing programs. In contrast, replicate B shows considerably greater stability. Yet, when examining batch accuracy - measured against the random training samples drawn each generation - performance became highly volatile across both replicates (see Figure 2).
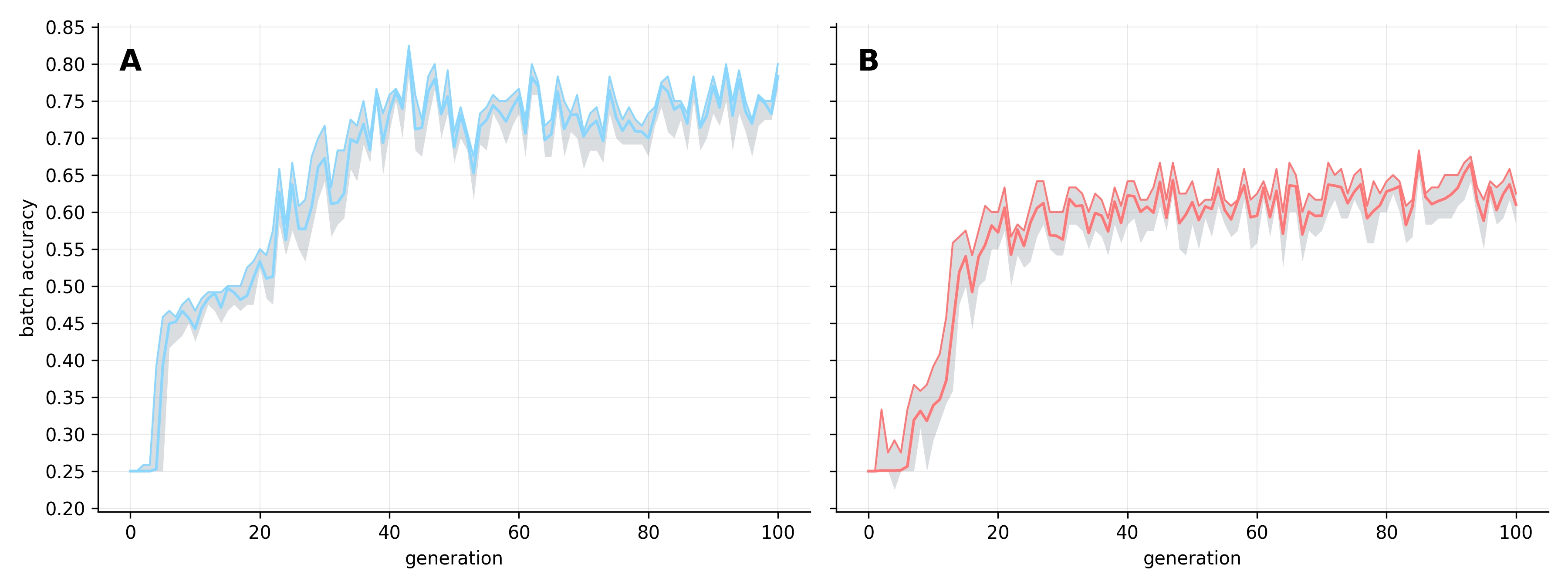
The high batch-level variability likely reflects sampling bias, where randomly drawn training batches may coincidentally favor specific programs and artificially inflate their batch-level accuracy. This concern is compounded by the narrow range of variation in batch accuracy, suggesting that selection pressure may be overly aggressive. This can lead to stochastic overfitting, where the population oscillates between accuracy levels as it chases the characteristics of individual batches. While this is a clear limitation for future research, it does not fully explain the intergenerational instability in test accuracy, which was largely unique to Replicate A.
3. Discussion
The experiments presented here represent some of our best results to date, reaching ~75% accuracy. Although this falls well short of state-of-the-art image classification methods, it illustrates how a broader paradigm - machine learning as algorithm discovery - can work in practice.
Traditional machine learning treats the problem as one of parameter estimation: given a model with a fixed, predetermined structure - a neural network, for instance - find the set of parameters, such as its weights, that best fits the data. We take a different view, framing the problem as one of algorithm discovery: we search for a sequence of rudimentary instructions - unconstrained by any particular structure - that can learn to make sense of data on its own terms.
For such a search to be truly open-ended, the primitive instructions being composed must be low-level enough to carry no implicit knowledge of the problem domain, while still including control flow operations that enable loops and conditional behavior. In principle, this leaves open the possibility that any algorithm capable of solving the given task could be discovered - including, conceivably, something resembling a neural network architecture.
This is, in essence, the aspiration of genetic programming: using the evolutionary process to search for algorithms, embodied as computer programs, that solve a given problem.
We pursue algorithm discovery by evolving Zyme programs. While built on an unconventional model of computation, they are composed entirely of low-level, byte-level instructions. Any operations on larger complex collections of input bytes - such as those needed for image processing - must be assembled autonomously from these primitive building blocks.
Achieving 75% accuracy on this subset of the MNIST image classification task therefore carries a precise meaning: through evolution alone, we have generated an algorithm that manipulates raw bytes and functions as an image classifier - without any implicit prior knowledge of the problem, without hand-crafted representations, and without any form of a priori feature extraction4. As far as we are aware, solving the problem in this way - discovering an algorithm composed of byte-level instructions - is without precedent.
The image processing algorithm that emerges from this process is evidently expressed in Zyme bytecode, since that's the evolvable representation used during discovery. But since the programs are ultimately composed of byte-level instructions, there is no theoretical barrier to extracting the underlying algorithm and transpiling it into other representations, better suited to different needs: such as Python for interpretability, or machine code for performance. The immediate implication is that extracted algorithms could be embedded directly into larger systems; but the more profound possibility lies in uncovering entirely new classes of algorithm. Stochastic or fuzzy algorithms are difficult for humans to reason about, and therefore remain underexplored and underrepresented in modern digital systems, yet these are precisely the kinds that nature routinely finds and exploits. Perhaps, algorithm discovery through evolution can offer a path toward unearthing such algorithms, especially those human intuition alone is unlikely to produce.
For now though, such possibilities remain distant. While 75% accuracy marks a symbolic milestone, it is still far from competitive with modern machine learning methods, and it is possible that this result is at or near the performance ceiling of the approach. That said, progress has been steady, and we have only just begun to explore the mechanisms available for improving and optimising performance. Here, we demonstrated gains from increasing the training dataset - still a fraction of the full MNIST set - and from fine-tuning a handful of population and evolutionary parameters, all while using relatively modest compute. Many avenues for optimisation remain entirely unexplored, including the core Zyme architecture itself: its instruction set and underlying design assumptions. The current instruction set was designed to be a minimal viable set: both familiar to humans and useful for debugging the system, rather than one optimised for evolvability. We have no immediate plans to revisit these design decisions, as other priorities take precedence for now, but they represent a significant source of untapped potential.
Taken together, these results and the breadth of unexplored optimisation suggest significant opportunity for improvement. We believe that continued progress can further demonstrate not only the underappreciated potential of genetic programming, but also the viability of the broader reframing it invites — machine learning as algorithm discovery.
1
In the previous experiments, the model was trained on a subset of the MNIST dataset comprising 2,080 image-label pairs (520 per category) and evaluated on 80 test pairs (20 per category). Here, we expanded the training set to 4,000 pairs (1,000 per category) and the test set to 200 pairs (50 per category). Alongside this, we increased the batch size used to evaluate candidate programs at each generation - the number of training examples sampled per category rose from 10 to 30.
We also substantially increased the number of candidate programs generated at each generation, improving the likelihood of discovering a beneficial mutation. We increased both the number of parent program pairs selected for recombination and the number of offspring each pair produces - rising from 500 to 2,000 mating pairs and from 30 to 63 offspring per pair. As a result, each generation now produces 126,000 new candidate programs, compared with 15,000 previously. Finally, the retained population size was doubled from 50 to 100.
2
In eukaryotic organisms, the vast majority of genomic DNA - approximately 98% - does not encode proteins. Initially, this non-coding DNA was dismissed as 'junk', assumed to be accumulated evolutionary debris given its lack of discernable function. We now know, however, that much of it serves a critical regulatory purpose, helping to govern the timing and manner in which genes are switched on and off. Importantly, because these regions are subject to far weaker evolutionary constraints than protein-coding sequences, they are free to accumulate mutations at a much higher rate, making them a potentially powerful engine for the emergence of new genes and biological functions.
3
Astute readers will note that, unlike our previous experiments, there are no longer individuals with 0% accuracy. You can see this in the previous Figure 2 the grey shaded area - representing the full range of population performance - frequently touches the baseline. We now recognize this is an artifact of our evaluation logic: if a program times out or fails to produce a valid output for a single input-output test pair, we immediately terminate its evaluation on further test pairs and assign a 0% score.
While this 'fail-fast' approach avoids wasting CPU cycles on programs already identified as faulty, it reveals an unusual challenge of evolving programs: that programs can fail to provide any output at all. Our programs must not only predict a label for a given input image, but provide that label in a strict format within a virtual CPU time limit. An individual program may be successful on the examples in the training batch, but fail to provide a valid output on the test set, triggering the 0% accuracy rating. We have yet to find a remedy for these viability mismatches between training and test data; even with larger batch sizes, one or two such programs can appear in a generation. For clarity, these outliers have been excluded from Figure 1 in this post.
4
One could argue that the design of the initial programs used to seed the starting population constitutes a form of prior information - and if so, this would undermine the claim of autonomous algorithm discovery. The concern is legitimate: it would certainly be possible to craft an initial program that encodes subtle hints about the nature of the input data, effectively smuggling in domain knowledge. We think of this as analogous to the choice of prior in Bayesian inference: some initial expectation is unavoidable, but a principled prior should be only weakly informative. In this experiment, the initial program is a crude hash function that encodes nothing about image processing and imposes no assumptions about the structure of the data. Whatever algorithm the evolved programs come to embody is, therefore, genuinely discovered - not inherited from the initial bespoke program.